Molecules exhibit non-reciprocal interactions without external forces, new study finds
Researchers from the University of Maine and Penn State discovered that molecules experience non-reciprocal interactions without external forces.
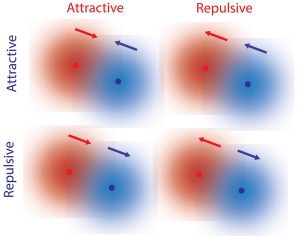
Fundamental forces such as gravity and electromagnetism are reciprocal, where two objects are attracted to each other or are repelled by each other. In our everyday experience, however, interactions don’t seem to follow this reciprocal law. For example, a predator is attracted to prey, but the prey tends to flee from the predator. Such non-reciprocal interactions are essential for complex behavior associated with living organisms. For microscopic systems such as bacteria, the mechanism of non-reciprocal interactions have been explained by hydrodynamic or other external forces, and it was previously thought that similar types of forces could explain interactions between single molecules.
In work published in the prestigious Cell Press journal Chem, UMaine theoretical physicist R. Dean Astumian and collaborators Ayusman Sen and Niladri Sekhar Mandal at Penn State have published a different mechanism by which single molecules can interact non-reciprocally without hydrodynamic effects. This mechanism invokes the local gradients of reactants and products due to the reactions facilitated by every chemical catalyst, a biological example of which is an enzyme. Because the response of a catalyst to the gradient depends on the catalyst’s properties, it is possible to have a situation in which one molecule is repelled by, but attracts, another molecule.
The authors’ “Eureka moment” occurred when, in their discussion, they realized that a property of every catalyst known as the kinetic asymmetry controls the direction of response to a concentration gradient. Because kinetic asymmetry is a property of the enzyme itself, it can undergo evolution and adaptation. The non-reciprocal interactions allowed by kinetic asymmetry also play a crucial role in allowing molecules to interact with each other, and may have played a critical role in the processes by which simple matter becomes complex.
Much previous work has been done by other researchers on what happens when non-reciprocal interactions occur. These efforts have played a central role in the development of a field known as “active matter.” In this earlier work, the non-reciprocal interactions were introduced by incorporation of ad hoc forces. The research described by Mandal, Sen and Astumian, however, describes a basic molecular mechanism by which such interactions can arise between single molecules. This research builds on earlier work in which the same authors showed how a single catalyst molecule could use energy from the reaction it catalyzed to undergo directional motion in a concentration gradient.
The kinetic asymmetry that features in determining the non-reciprocal interactions between different catalysts has also been shown to be important for the directionality of biomolecular machines, and has been incorporated in the design of synthetic molecular motors and pumps.
The collaboration between Astumian, Sen and Mandal aims to reveal the organizational principles behind loose associations of different catalysts that may have formed the earliest metabolic structures that eventually led to the evolution of life.
“We’re at the very beginning stages of this work, but I see understanding kinetic asymmetry as a possible opportunity for understanding how life evolved from simple molecules,” Astumian says. “Not only can it provide insight into complexification of matter, kinetic asymmetry can also be used in the design of molecular machines and associated technologies.”
Astumian joined UMaine’s Department of Physics and Astronomy in 2001. His research focuses on biophysics, condensed matter physics, and chemically driven molecular machines.
He was named a fellow of the American Association for the Advancement of Science (AAAS) In 2016. His other honors include the Galvani Prize of the Bio-electrochemical Society, the Humboldt Prize, the Feynman Prize, and the Royal Society of Chemistry Horizon Prize, the Perkin prize in physical organic chemistry.
Contact: Marcus Wolf, 207.581.3721; marcus.wolf@maine.edu
